José Luis Oliveira
Universidade de Aveiro, DETI / IEETA
3810-193 Aveiro, Portugal
jlo@ua.pt
(+351) 234 370 500
ANACONDA is a software package specially developed for the study of genes’ primary structure. It uses gene sequences downloaded from public databases, as FASTA and GenBank, and it applies a set of statistical and visualization methods in different ways, to reveal information about codon context, codon usage, nucleotide repeats within open reading frames (ORFeome) and others.
Codon context analysis
Genome sequencing is opening unprecedent ways for understanding how gene primary structure is organized. Two of the most studied open reading frame characteristics are codon usage and codon context.
Traditional methods used for codon usage and context analysis do not provide user-friendly tools to carry out detailed gene primary structure analysis at a genomic scale.
Codon usage tables, using absolute metric, are available in public databases for any sequenced gene or genome and freeware software for multivariate analysis (correspondence analysis) of codon and amino acid usage is also readily available, however sophisticated statistical and data visualization tools are clearly lacking.
We propose the usage of several statistical methods – contingency table analysis, residual analysis, multivariate analysis (cluster analysis) – to analyze the codon bias under various aspects (degree of association, contexts and clustering).
Cluster analysis
A cluster analysis tool allows also calculating similarities between two vectors of the contingency table. This technique is used to group lines and columns (codons) of the correlation matrix, allowing highlight global patterns in the genes.
The statistical tools that are incorporated in the system, for data clustering, residual analysis and histogram plotting of calculated indexes, allow reaching new conclusions on gene primary structure features at a genomic scale. We expect that the results obtained will permit identifying some general rules that govern codon context and codon usage in any genome. Additionally, the identification of genes containing expanded codons that arise as a consequence of erroneous DNA replications events will permit uncovering new genes associated to human disease.
Visualization
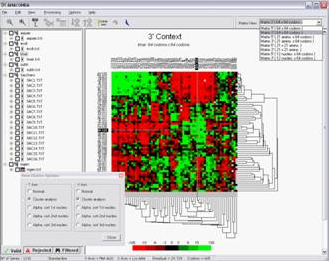
In order to detect the impact of codon context bias (as well as the presence of rare codons) on coding sequences, ANACONDA has additional tools for sequence mapping. The layout for sequence include written information about the ORF and the sequence itself, in which the codons have been coloured with the same residual colour scale of the ORFeome map.
ANACONDA allows the user to work with more than one ORFeome at a time. This creates large data sets that are difficult to deal with, in particular when multiple comparisons are being performed.
Considering that vast number of ORFeomes can be analyzed simultaneously by ANACONDA, we have included extra tools to allow comparative studies.
he statistical tools that are incorporated in the system, for data clustering, residual analysis and histogram plotting of calculated indexes, allow reaching new conclusions on gene primary structure features at a genomic scale. We expect that the results obtained will permit identifying some general rules that govern codon context and codon usage in any genome.
Past publications
- G. Moura, M. Pinheiro, J. Arrais, A. C. Gomes, L. Carreto, A. Freitas, J. L. Oliveira, and M. A. Santos, “Large Scale Comparative Codon-Pair Context Analysis Unveils General Rules that Fine-Tune Evolution of mRNA Primary Structure”, PLoS ONE, vol. 2, no. 9, e847, doi:10.1371/journal.pone.0000847, 2007.
- M. Pinheiro, V. Afreixo, G. Moura, A. Freitas, M. A. Santos, and J. L. Oliveira, “Statistical, computational and visualization methodologies to unveil gene primary structure features”, Methods of Information in Medicine, vol. 45, no. 2, pp. 163-168, 2006.
- G. Moura, M. Pinheiro, R. Silva, I. Miranda, V. Afreixo, G. Dias, A. Freitas, J. L. Oliveira, and M. A. Santos, “Comparative context analysis of codon pairs on an ORFeome scale”, Genome Biology, vol. 6, no. 3, pp. R28, 2005.
Download
Anaconda 2 is now available for download. It is freely available for fundamental research only.
[Download]
Last Update (2011-01-12)
New features (Version 2.0, 2011):
- Corrected some bugs
- Enriched codon statistics and visualization maps
- Comparing context maps across several species
- Integration of tRNA copy number processing
- Single and multiple sequence alignments using codon context (BLASTP and ClustalW)